On this article, you’ll learn to select between PCA and t-SNE for visualizing high-dimensional information, with clear trade-offs, caveats, and dealing Python examples.
Subjects we are going to cowl embody:
- The core concepts, strengths, and limits of PCA versus t-SNE.
- When to make use of every methodology — and when to mix them.
- A sensible PCA → t-SNE workflow with scikit-learn code.
Let’s not waste any extra time.
Selecting Between PCA and t-SNE for Visualization (click on to enlarge)
Picture by Editor
For information scientists, working with high-dimensional information is a part of every day life. From buyer options in analytics to pixel values in photos and phrase vectors in NLP, datasets usually comprise a whole lot and hundreds of variables. Visualizing such complicated information is tough.
That’s the place dimensionality discount strategies are available in. Two of probably the most broadly used strategies are Principal Element Evaluation (PCA) and t-Distributed Stochastic Neighbor Embedding (t-SNE). Whereas each cut back dimensions, they serve very totally different objectives.
Understanding Principal Element Evaluation (PCA)
Principal Element Evaluation is a linear methodology that transforms information into new axes known as principal elements. Its aim is to transform your information into a brand new coordinate system the place the best variations lie on the primary axis (the primary principal part), the second best on the second axis, and so forth. It does this by performing an eigendecomposition (the method of breaking down a sq. matrix into a less complicated, “canonical” kind utilizing its eigenvalues and eigenvectors) of the information covariance matrix or a Singular Worth Decomposition (SVD) of the information matrix.
These elements seize the very best variance within the information and are ordered from most necessary to least necessary. Consider PCA as rotating your dataset to search out one of the best angle that exhibits probably the most unfold of data.
Key Benefits and When to Use PCA
- Characteristic Discount & Preprocessing: Use PCA to cut back the variety of enter options for a downstream mannequin (like regression or classification) whereas retaining probably the most informative indicators.
- Noise Discount: By discarding elements with minor variance (usually noise), PCA can clear your information.
- Interpretable Elements: You possibly can examine the
components_attribute to see which authentic options contribute most to every principal part. - World Variance Preservation: It faithfully maintains large-scale distances and relationships in your information.
Implementing PCA with Scikit-Be taught
Utilizing PCA in Python’s scikit-learn is easy. The important thing parameter is n_components, which defines the variety of dimensions to your output.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 |
from sklearn.decomposition import PCA from sklearn.datasets import load_iris import matplotlib.pyplot as plt
# Load pattern information iris = load_iris() X = iris.information y = iris.goal
# Apply PCA, lowering to 2 dimensions for visualization pca = PCA(n_components=2) X_pca = pca.fit_transform(X)
# Visualize the outcome plt.determine(figsize=(8, 6)) scatter = plt.scatter(X_pca[:, 0], X_pca[:, 1], c=y, cmap=‘viridis’, edgecolor=‘ok’, s=70) plt.xlabel(‘Principal Element 1’) plt.ylabel(‘Principal Element 2’) plt.title(‘PCA of Iris Dataset’) plt.colorbar(scatter, label=‘Iris Species’) plt.present()
# Look at defined variance print(f“Variance defined by every part: {pca.explained_variance_ratio_}”) print(f“Complete variance captured: {sum(pca.explained_variance_ratio_):.2%}”) |
This code reduces the four-dimensional Iris dataset to 2 dimensions. The ensuing scatter plot exhibits the information unfold alongside axes of most variance, and the explained_variance_ratio_ tells you the way a lot info was preserved.
Code output:
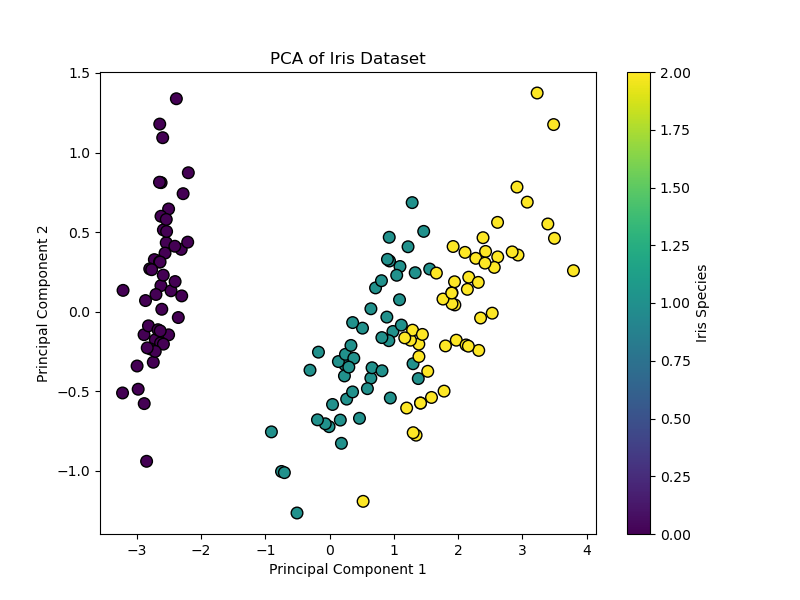
|
Variance defined by every part: [0.92461872 0.05306648] Complete variance captured: 97.77% |
When to Use PCA
- If you wish to cut back options earlier than machine studying fashions
- If you wish to take away noise
- If you wish to velocity up coaching
- If you wish to perceive world patterns
Understanding t-Distributed Stochastic Neighbor Embedding (t-SNE)
t-SNE is a non-linear method designed virtually totally for visualization. It really works by modeling pairwise similarities between factors within the high-dimensional area after which discovering a low-dimensional (2D or 3D) illustration the place these similarities are finest maintained. It’s notably good at revealing native constructions like clusters that could be hidden in excessive dimensions.
Key Benefits and When to Use t-SNE
- Visualizing Clusters: It’s nice for creating intuitive, cluster-rich plots from complicated information like phrase embeddings, gene expression information, or photos
- Revealing Non-Linear Manifolds: It will probably reveal detailed, curved constructions that linear strategies like PCA can not
- Give attention to Native Relationships: Its design ensures that factors shut within the authentic area stay shut within the embedding
Important Limitations
- Axes Are Not Interpretable: The t-SNE plot’s axes (t-SNE1, t-SNE2) don’t have any elementary which means. Solely the relative distances and clustering of factors are informative
- Do Not Examine Clusters Throughout Plots: The size and distances between clusters in a single t-SNE plot are usually not similar to these in one other plot from a special run or dataset
- Perplexity is Key: That is a very powerful parameter. It balances the eye between native and world construction (typical vary: 5–50). You could experiment with it
Implementing t-SNE with Scikit-Be taught
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 |
from sklearn.datasets import load_iris from sklearn.manifold import TSNE import matplotlib.pyplot as plt
# Load pattern information iris = load_iris() X = iris.information y = iris.goal
# Apply t-SNE. Word the important thing ‘perplexity’ parameter. tsne = TSNE(n_components=2, perplexity=30, random_state=42, init=‘pca’) X_tsne = tsne.fit_transform(X)
# Visualize the outcome plt.determine(figsize=(8, 6)) scatter = plt.scatter(X_tsne[:, 0], X_tsne[:, 1], c=y, cmap=‘viridis’, edgecolor=‘ok’, s=70) plt.xlabel(‘t-SNE Element 1 (no intrinsic which means)’) plt.ylabel(‘t-SNE Element 2 (no intrinsic which means)’) plt.title(‘t-SNE of Iris Dataset (Perplexity=30)’) plt.colorbar(scatter, label=‘Iris Species’) plt.present() |
This code creates a t-SNE visualization. Setting init="pca" (the default) makes use of a PCA initialization for higher stability. Discover the axes are intentionally labeled as having no intrinsic which means.
Output:
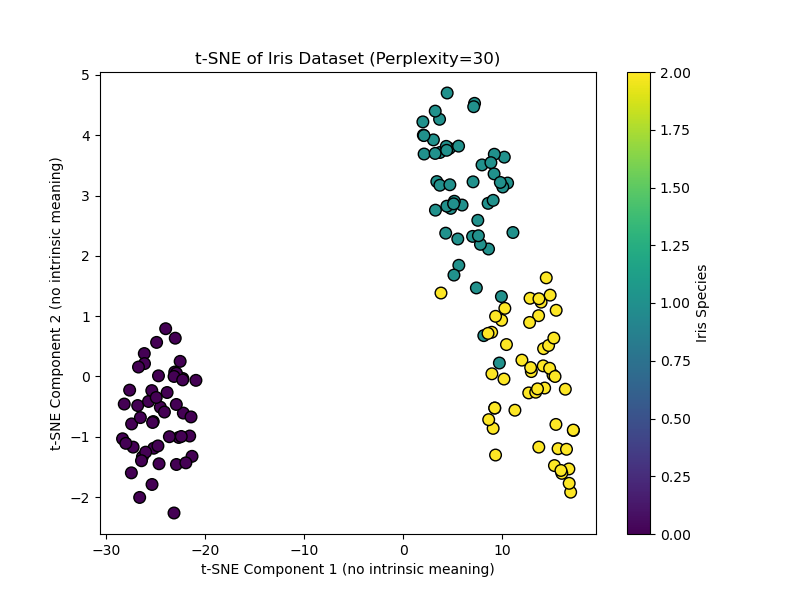
When to Use t-SNE
- If you wish to discover clusters
- When you could visualize embeddings
- If you wish to reveal hidden patterns
- It isn’t for characteristic engineering
A Sensible Workflow
A strong and customary finest follow is to mix PCA and t-SNE. This makes use of the strengths of each:
- First, use PCA to cut back very high-dimensional information (e.g., 1000+ options) to an intermediate variety of dimensions (e.g., 50). This removes noise and drastically hurries up the following t-SNE computation
- Then, apply t-SNE to the PCA output to get your closing 2D visualization
Hybrid strategy: PCA adopted by t-SNE
|
from sklearn.decomposition import PCA
# Step 1: Scale back to 50 dimensions with PCA pca_for_tsne = PCA(n_components=50) X_pca_reduced = pca_for_tsne.fit_transform(X_high_dim) # Assume X_high_dim is your authentic information
# Step 2: Apply t-SNE to the PCA-reduced information X_tsne_final = TSNE(n_components=2, perplexity=40, random_state=42).fit_transform(X_pca_reduced) |
The instance above demonstrates utilizing t-SNE to cut back to 2D for visualization, and the way PCA preprocessing could make t-SNE quicker and extra secure.
Conclusion
Selecting the best software boils all the way down to your major goal:
- Use PCA whenever you want an environment friendly, deterministic, and interpretable methodology for general-purpose dimensionality discount, characteristic extraction, or as a preprocessing step for an additional mannequin. It’s your go-to for a primary have a look at world information construction.
- Use t-SNE when your aim is only visible exploration and cluster discovery in complicated, non-linear information. Be ready to tune parameters and by no means interpret the plot quantitatively
Begin with PCA. If it reveals clear linear developments, it could be ample. In case you suspect hidden clusters, change to t-SNE (or use the hybrid strategy) to disclose them.
Lastly, whereas PCA and t-SNE are foundational, pay attention to fashionable alternate options like Uniform Manifold Approximation and Projection (UMAP). UMAP is usually quicker than t-SNE and is designed to protect extra of the worldwide construction whereas nonetheless capturing native particulars. It has turn out to be a well-liked default selection for a lot of visualization duties, persevering with the evolution of how we see our information.
I hope this text gives a transparent framework for selecting between PCA and t-SNE. The easiest way to construct this understanding is to experiment with each strategies on datasets you already know effectively, observing how their totally different natures form the story your information tells.

